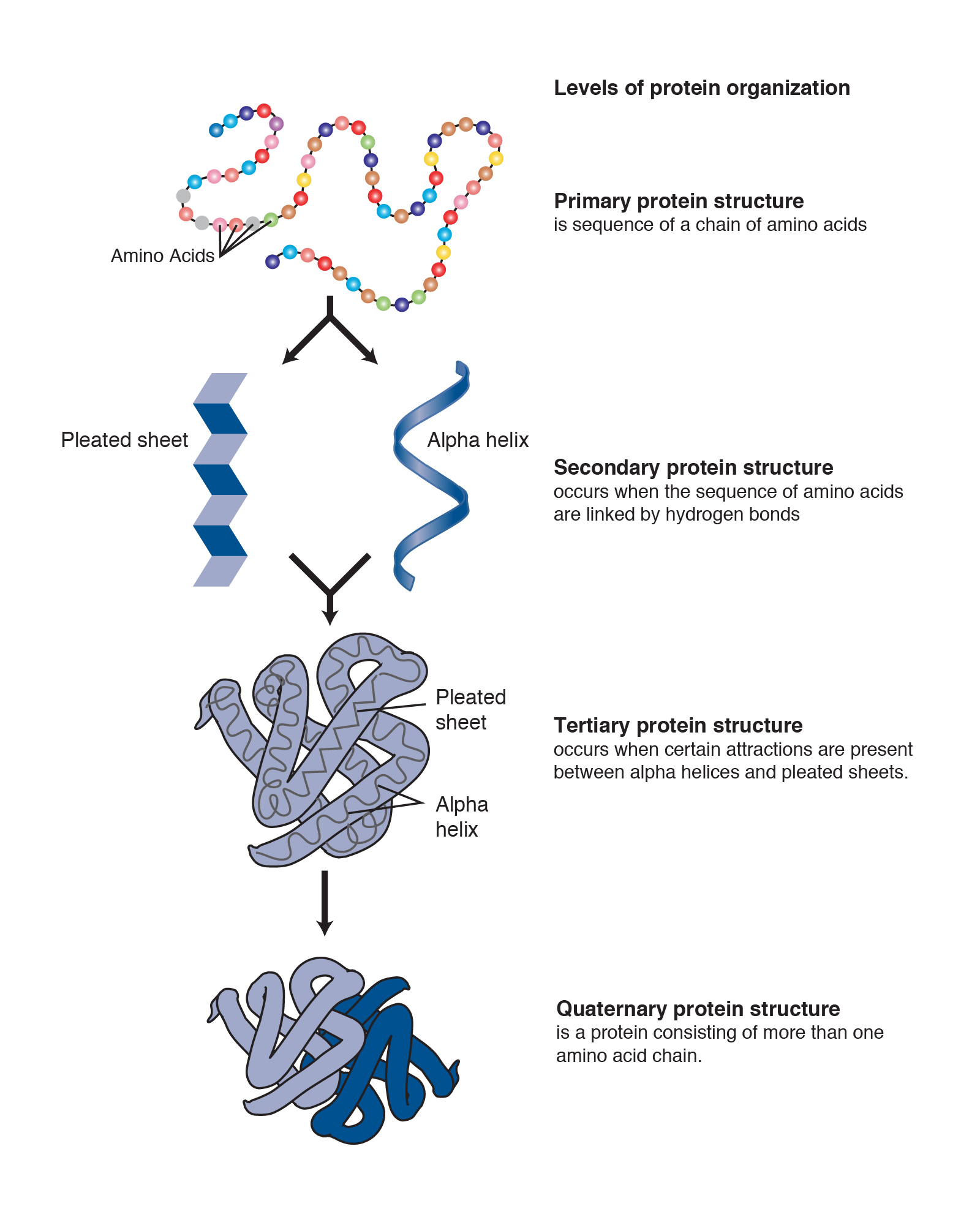
The different R-groups have different characteristics based on the nature of atoms incorporated into the functional groups. There are a total of 20 alpha amino acids that are commonly incorporated into protein structures (Figure 2.x). The basic structure of an amino acid is shown below:įigure 2.1 General Structure of an Alpha Amino Acid 
They differ from one another only at the R-group position. Within living organisms there are 20 amino acids used as protein building blocks. In the diagram below, this group is designated as an R-group. In addition to the amine and the carboxylic acid, the alpha carbon is also attached to a hydrogen and one additional group that can vary in size and length. The alpha designation is used to indicate that these two functional groups are separated from one another by one carbon group. As their name implies they contain a carboxylic acid functional group and an amine functional group. The major building block of proteins are called alpha (α) amino acids. They are all, however, polymers of alpha amino acids, arranged in a linear sequence and connected together by covalent bonds. Their structures, like their functions, vary greatly. Each cell in a living system may contain thousands of different proteins, each with a unique function. 
Proteins may be structural, regulatory, contractile, or protective they may serve in transport, storage, or membranes or they may be toxins or enzymes. Proteins are one of the most abundant organic molecules in living systems and have the most diverse range of functions of all macromolecules. Adapted with permission from Applied Biosystems Procise User’s Manual.Chapter 2: Protein Structure 2.1 Amino Acid Structure and Properties 2.2 Peptide Bond Formation and Primary Protein Structure 2.3 Secondary Protein Structure 2.4 Supersecondary Structure and Protein MotifsĢ.5 Tertiary and Quaternary Protein Structure 2.6 Protein Folding, Denaturation and Hydrolysis 2.7 References The HPLC pumps and detector are controlled through the sequencer computer during the sequencing process in the automated mode. The HPLC injector transfers the PTH amino acids from the conversion flask to an HPLC column. Delivery of appropriate chemicals or solvents to the sample cartridges or conversion flask are regulated by the related reagent or solvent valve blocks. The conversion reactions section includes: aqueous acid (R4) for conversion of the amino acid derivative to PTH amino acids acetonitrile/water (S4) to redissolve the PTH amino acids for HPLC analysis PTH standard (R5) used for calibration and additional positions for user-designated chemistries or solvents (X2 and X3). Bottles for chemistry involved in sequencer reactions include: base (R2) and PITC (R1) trifluoroacetic acid (R3) for cleavage, which can be delivered in either the gas phase or as a small pulse of liquid reagent solvents for removing reaction byproducts (S1 and S2) and extraction of the cleaved amino acid derivative to the conversion flask (S3) and additional reagent positions for addition of optional chemistries or solvents (X1 and X3). Finally, discussion of optimization of sequencer performance as well as possible solutions to frequently encountered problems is included.Īpplied Biosystems Procise 494 reagent schematic, illustrating the current complexity of instrumentation used for automated sequence analysis. A discussion of data interpretation is therefore provided. The amount of data obtained from a single sequencer run is substantial, and careful interpretation of this data by an experienced scientist familiar with the current operation performance of the instrument used for this analysis is critically important. Methods are provided for optimizing separation of PTH amino acid derivatives on Perkin-Elmer instruments and for increasing the proportion of sample injected onto the PTH analyzer on older Perkin-Elmer instruments by installing a modified sample loop. 
Sequence analysis of protein or peptide samples in solution or bound to PVDF membranes using a Hewlett-Packard Model G1005A sequencer is also described. This unit describes the sequence analysis of protein or peptide samples in solution or bound to PVDF membranes using a Perkin-Elmer Procise Sequencer. Amino-terminal (N-terminal) sequence analysis is used to identify the order of amino acids of proteins or peptides, starting at their N-terminal end.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
